iDPGK is a web server for identifying phosphoglycerylation sites. Protein phosphoglycerylation is the reaction in which a 1,3-bisphosphoglyceric acid (1,3-BPG) is attached to a lysine residue of a protein molecule, and thus to form a 3-phosphoglyceryl-lysine (pgK). Phosphoglycerylation is a reversible and non-enzymatic post-translational modification (PTM). It has been shown that the process of lysine phosphoglycerylation plays important regulatory roles in glucose metabolism and glycolytic process. However, studies in this field have been limited by the difficulty of experimentally identifying modified site specificity in lysine phosphoglycerylation (PGK).
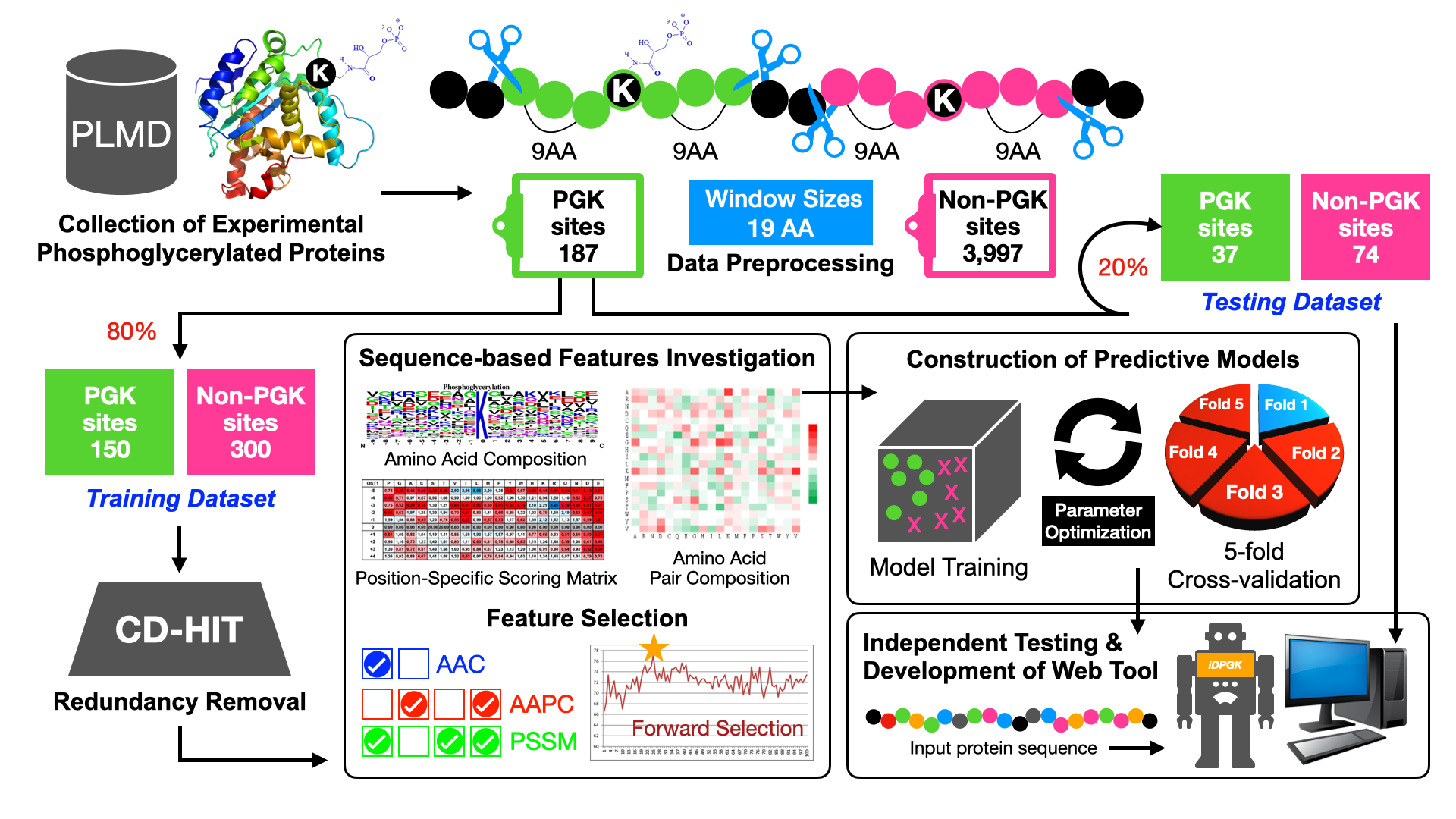
To facilitate this process, several tools have been proposed for the computational identification of PGK sites. In this study, we developed a new method to investigate the specificities of lysine succinylation sites based on the sequence-based features. With the experimentally verified PGK sites collected from public resources, the entropy plots of sequence logo reveal the conserved motifs in modified sites to which a 1,3-BPG can be attached. The libsvm algorithm was then applied to train a PGK site prediction model with not only the amino acid composition (AAC) feature but also the amino acid pair composition (AAPC), BLOSUM62 matrix (B62) and position-specific scoring matrix (PSSM). Evaluation using the 5-fold cross validation approach showed that the predictive model trained with the selected features significantly outperformed existing tools. The effectiveness of the constructed model was further evaluated with experimentally verified lysine phosphoglycerylation sites manually extracted from published research articles.